Agricultural Science and Food Processing | Volume 2, Issue 1: 47-55, 2025 | DOI: 10.62762/ASFP.2024.865235
Abstract
Inverse PCR (IPCR) is a reliable, straightforward, and effective technique for acquiring unknown sequences. In this study, we used the model monocot Brachypodium distachyon (ecotype Bd21) to standardize the conditions and materials necessary for the successful execution of IPCR. The analysis of the amplified sequences resulted in the following conclusions. First, the distance between the nearest primer and the boundary of the known-unknown sequence is crucial for determining whether the target sequence can be expanded in the second round of IPCR. Specifically, this distance should exceed 100 bp, ideally around 200 bp. Second, because the random cleavage of a 6 bp endonuclease occurs at a gre... More >
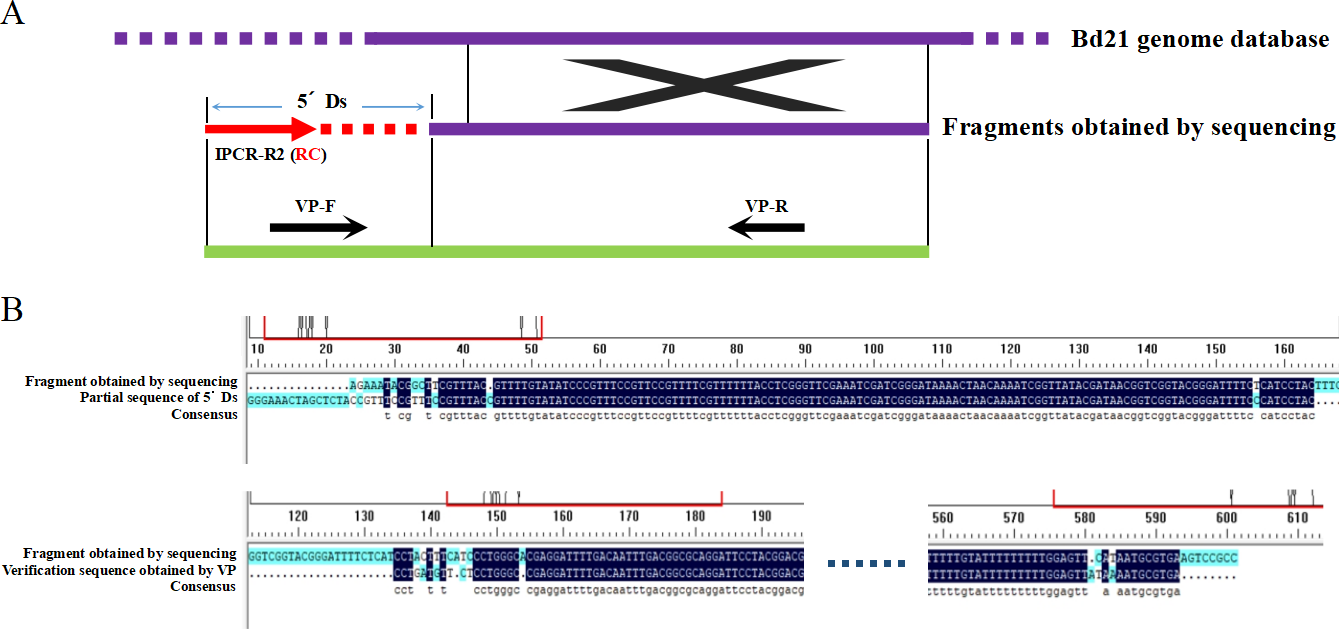
Graphical Abstract